 The Data Mining Forum
The Data Mining Forum 



IMPORTANT: This is the old Data Mining forum.
I keep it online so that you can read the old messages.
Please post your new messages in the new forum: https://forum2.philippe-fournier-viger.com/index.php
Discussed the execution time efficiency of the Frequent Sequence Pattern mining process.
Posted by:
Long Pham
Date: April 07, 2014 04:04AM
HI All.
Finding all Frequent Sequence Pattern is a challenging and time-consuming problem due to reason.
- it may be hard to find all Frequent Sequence Pattern in dense and very large datasets
All algorithms have been implemented by sequential strategies. their runtime performances have not been satisfied on large datasets, especially recently emerged dense datasets.Researchers have begun switching to parallel and distributed computing techniques to accelerate the computation.
How to implement parallel Algorithms to Finding all frequent sequence pattern ? Can you help me to make a good parallel algorithms.
I am very appreciated for your help.
Finding all Frequent Sequence Pattern is a challenging and time-consuming problem due to reason.
- it may be hard to find all Frequent Sequence Pattern in dense and very large datasets
All algorithms have been implemented by sequential strategies. their runtime performances have not been satisfied on large datasets, especially recently emerged dense datasets.Researchers have begun switching to parallel and distributed computing techniques to accelerate the computation.
How to implement parallel Algorithms to Finding all frequent sequence pattern ? Can you help me to make a good parallel algorithms.
I am very appreciated for your help.
Re: Discussed the execution time efficiency of the Frequent Sequence Pattern mining process.
Posted by:
webmasterphilfv
Date: April 07, 2014 05:37AM
There is at least two ways to handle very large datasets:
- perform sampling (use only a subset of the database). For example, even if there are millions of sequences, you may just need 10,000 to get enough patterns that are useful and representative.
- use some parallel algorithm. Several sequential pattern mining algorithm can be parallelized easily. For example, the SPADE and SPAM algorithm can be easily parallelized because they are divide and conquer algorithms. In SPMF, there is a multi-thread version of SPADE available in the source code as an example. It will work in parallel for a multi-core processor.
- perform sampling (use only a subset of the database). For example, even if there are millions of sequences, you may just need 10,000 to get enough patterns that are useful and representative.
- use some parallel algorithm. Several sequential pattern mining algorithm can be parallelized easily. For example, the SPADE and SPAM algorithm can be easily parallelized because they are divide and conquer algorithms. In SPMF, there is a multi-thread version of SPADE available in the source code as an example. It will work in parallel for a multi-core processor.
Re: Discussed the execution time efficiency of the Frequent Sequence Pattern mining process.
Posted by:
Long Pham
Date: April 10, 2014 09:44PM
I would like to have the PRISM algorithm in C# or Java.
Is Prism algorithm the best Algorithm to Mining Frequent Sequence Pattern?
Thanks.
Is Prism algorithm the best Algorithm to Mining Frequent Sequence Pattern?
Thanks.
Re: Discussed the execution time efficiency of the Frequent Sequence Pattern mining process.
Posted by:
webmasterphilfv
Date: April 11, 2014 09:34AM
I'm not sure that Prism is so fast. It was compared with Spade, PrefixSpan and SPAM.
It was not compared with more recent algorithms such as BitSPADE which is supposed to be faster than SPADE for example.
You may have a look at my algorithm CM-SPADE and CM-SPAM in the open-source of the SPMF library in Java. CM-SPADE is generally much faster than SPADE (and our implementation of SPADE similar to BitSPADE), SPAM, PrefixSpan, GSP, etc. You can see a graph here:
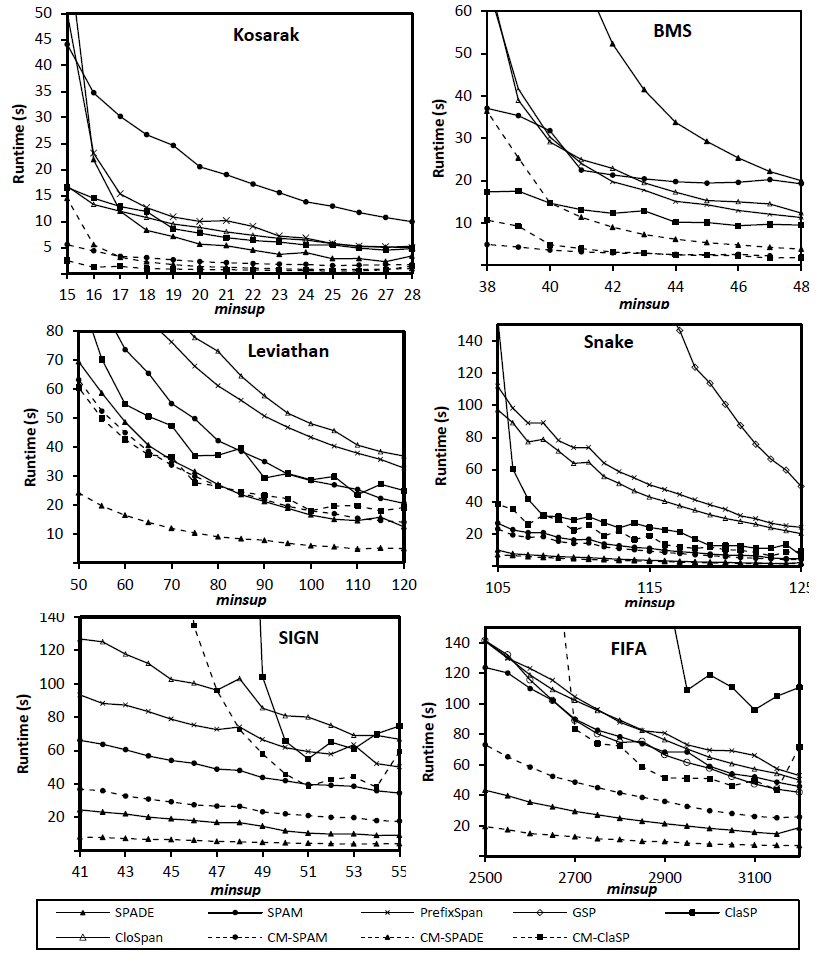
The paper about CM-SPADE/CM-SPAM will be published at PAKDD2014 next month. But the source code is already available.
There are also some other algorithms that have been proposed but the problem is always that every authors do not always compare their work with every other works. Thereore, it is not always clear which one is the best.
Edited 1 time(s). Last edit at 04/11/2014 09:35AM by webmasterphilfv.
It was not compared with more recent algorithms such as BitSPADE which is supposed to be faster than SPADE for example.
You may have a look at my algorithm CM-SPADE and CM-SPAM in the open-source of the SPMF library in Java. CM-SPADE is generally much faster than SPADE (and our implementation of SPADE similar to BitSPADE), SPAM, PrefixSpan, GSP, etc. You can see a graph here:

The paper about CM-SPADE/CM-SPAM will be published at PAKDD2014 next month. But the source code is already available.
There are also some other algorithms that have been proposed but the problem is always that every authors do not always compare their work with every other works. Thereore, it is not always clear which one is the best.
Edited 1 time(s). Last edit at 04/11/2014 09:35AM by webmasterphilfv.
Re: Discussed the execution time efficiency of the Frequent Sequence Pattern mining process.
Posted by:
Long Pham
Date: April 11, 2014 05:04PM
Hi Philippe,
Thanks for the quick answer.
Thanks for the quick answer.
Re: Discussed the execution time efficiency of the Frequent Sequence Pattern mining process.
Posted by:
Dang Nguyen
Date: April 13, 2014 06:40PM
Hi Long,
You can adapt the ideas from two below papers for parallel mining sequence patterns.
1. An effective parallel approach for genetic-fuzzy data mining
http://www.sciencedirect.com/science/article/pii/S0957417413005915
2. Efficient strategies for parallel mining class association rules
http://www.sciencedirect.com/science/article/pii/S0957417414000621
The first paper implements a parallel algorithm on the Master-Slave model. The second one utilizes the multi-core processor architecture.
Best,
Dang Nguyen
You can adapt the ideas from two below papers for parallel mining sequence patterns.
1. An effective parallel approach for genetic-fuzzy data mining
http://www.sciencedirect.com/science/article/pii/S0957417413005915
2. Efficient strategies for parallel mining class association rules
http://www.sciencedirect.com/science/article/pii/S0957417414000621
The first paper implements a parallel algorithm on the Master-Slave model. The second one utilizes the multi-core processor architecture.
Best,
Dang Nguyen
Re: Discussed the execution time efficiency of the Frequent Sequence Pattern mining process.
Posted by:
Long Pham
Date: April 16, 2014 06:15PM
hi Dang,
Thank you for helping
Thank you for helping